Resources
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Expand
Collapse
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group

Combined TIRFM and AFM
Dr. Atom Sarkar, now a neurosurgeon at Ohio State University Medical Centre (Columbus, Ohio), in collaboration with Prof Julio Ferdandez from Columbia University (New York, NY), has been interested in the role of nanotechnology in neurosurgery. Currently he is delving into the world of protein mechanics. Why? Because a knowledge of a protein's structural dynamics is central to the understanding of many neurological diseases ranging from brain tumors to neurodegenerative processes such as Alzheimer's Disease. Protein folding and unfolding happen on the nanometer scale. Therefore Atom needs a technique of nanometer or sub-nanometer resolution. Confocal won't cut it. He has been using a customized total internal reflection (TIRF)/atomic force microscope (AFM)/electron multiplying charge coupled device (EMCCD) to study protein unfolding, with the aim of developing a TIRF-only method to follow protein dynamics with sub-nanometer resolution. This will open protein unfolding experiments to many more scientists.
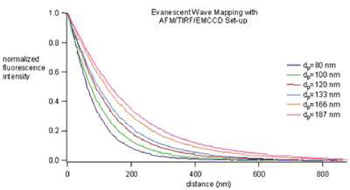
Measurement of a series of evanescent penetration depths
The key principle behind the methodology is the use of a nanometer-scale calibrated evanescent wave to measure the position of a fluorophore moving along the optical axis. By using a TIRFM generated evanescent wave that has an intensity which decays exponentially as a function of vertical distance, Atom can correlate changes from fluorescence intensity into displacements in the vertical axis. In order to calibrate the evanescent wave, an AFM/TIRFM/EMCCD device was constructed consisting of an AFM head mounted on top of a TIRFM microscope equipped with an electron multiplying charge coupled device (EMCCD) (see Combined TIRFM and AFM figure). The system exploits the distance dependent evanescent wave as a "ruler" to deconvolve fluorescent intensity into length (see figure above).
The six curves depicted above nicely illustrate the measured evanescent field decay and how by changing the TIRFM conditions, the evanescent wave penetration depth, dp, can be adjusted to fit experimental needs. The ability to measure distance with the evanescent field has been dubbed Evanescent Nanometry.
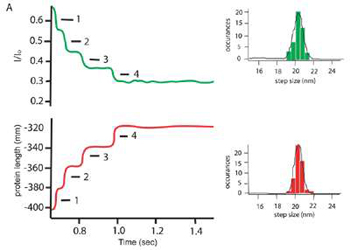
Evanescent Nanometry has already been used to follow the forced mediated unfolding of the protein ubiquitin, and these results rival the precision and accuracy obtained by standard AFM only studies.
Current experiments involve replacing the AFM entirely and attaching magnetic fluorescent beads to the proteins.
The magnetic beads allow the tethered proteins to be manipulated by an electromagnetic tweezer set-up, while the fluorescence of the bead enable nanometer precision monitoring of the proteins unfolding and folding dynamics, which are captured by the EMCCD.
